Chemoinformatics and Imaging
Background
Our work group aims at providing IT based solutions for biochemical aspects of the systems biology and systems medicine projects.
We develop and adapt software solutions to provide the required tools for medical research.
We accomplish this by applying and extending bioinformatics, cheminformatics and image processing methods to complement systems biology research questions integrating the means for statistical analysis. While extending our methodology toolbox we explicitly try to strengthen the planed analysis by integrating early on the requirements for the statistical analysis of the output of our pipelines.

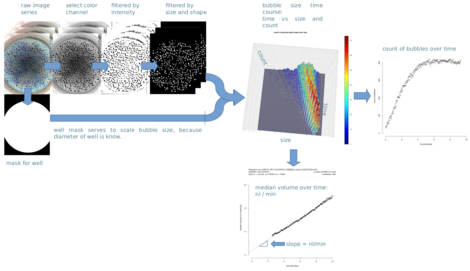
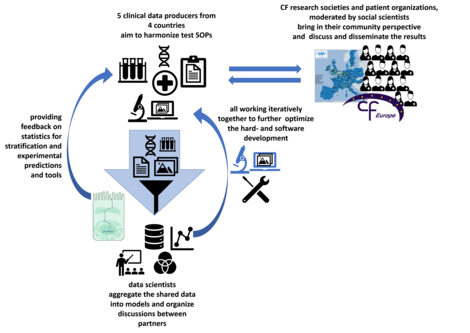
As more and more image based research is conducted and the number of created images per project increases, we develop the solutions for an automated analysis, e.g. AutoBuSTeD project, but if necessary can even venture out to help improve the acquisition setup. Our newest project Stracyfic is aiming to bring the diagnostic test to the clinical routine by developing the hardware and software for this diagnostic test.
Research foci
For the past years we focus our research in the field of cystic fibrosis, where we still see an unmet demand for IT solutions to help this field.
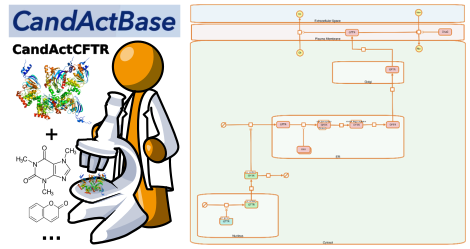
Our CandActCFTR project aims at providing means to distil and curate compound libraries from literature and other online sources and combine this structured information for analysis.
CF is a good target for a system medicine approach
At a first glance, CF is a monocausal disease in which over 2000 putative mutations leading to various forms of phenotypes have been identified. Among these, about 300 variants define the more common types. The CFTR protein is only effective as an integral membrane protein, and as such, it is affected by transcription, translation, folding and degradation, as well as protein traficking processes. Thus, this monogenic disease has multiple sites for potential drug intervention during its life cycle and covers also protein structure variants. This makes it an interesting target for a system medicine approach. The life cycle of the protein offers also multiple modes to obtain information to annotate the system and the existing literature offers various annotations for specific combinations of mutations and read-outs (e.g. protein expression, functional patch clamp measurements, up to structure models and molecular dynamic simulations).
Our newest additon - the CF-Map project - is a MINERVA-based implementation of the CFTR-lifecyle, which provides an interactive view of the underlying systems biology model. It is also the site where we showcase the integration of a specfic compound database like CandActCFTR with a systems biology model and will be extended with more modules in the future.
Web resources
- CandActCFTR a CFTR protein specific curated compound data base
- CF-disease map an interactive systems medicine model, with a focus on CFTR protein maturation
Current projects
Members of the group
Current members of the group
Dr. Manuel M. Nietert (Head of the group)
Former members of the group
- Tim Patrick Kakucs, Bachelor: Extracting structural parameters of the lung from 3D High-res Micro computed tomography (2024)
- Jonathan Tobias Hock, Bachelor: Automatisierte Generierung von DiseaseMaps: Ein Text Mining-Ansatz zur Beschleunigung der Krankheitsmodellierung (2024)
- Stephan Alves Dias, Master: Entwicklung einer Webapplikation zur lokalen Auswertung der bildgebenden Analyse zur Mukoviszidose-Diagnostik (HAWK/Göttingen) (2024)
- Dr. Liza Vinhoven, PhD: Cystic Fibrosis as a model use case for implementing cell based disease models in systems medicine(2022)
- Malte Voskamp, Master: Using text mining to annotate and construct biological disease maps (2021)
- David Tschritter, Bachelor: Automatische Quantifizierung von Lysosomenparametern aus Mikroskopiebildern im Kontext der zystischen Fibrose (Hochschule Emden/Leer) (2020)
- Paul Würzburg, Bachelor: Automated Image Analysis by Tracing Object Evolution in Time Series Data for a Diagnostic Test for Cystic Fibrosis (2020), Master: Training a Neural Network for Automatic Segmentation of Renal Masses in Computed Tomography Images with Uncertainty Estimation (2024)

contact information
- telephone: +49 551 3961789
- e-mail address: manuel.nietert(at)med.uni-goettingen.de
- location: 2.115
Publications
- Zeng, R.; Al-Bourini, O.; Lettermann, L.; Lettermann, L.; Olgemöller, U.; Hofer, S.; Boentert, M.; Friede, T.; Nietert, M.; Voit, D.; Frahm, J.; Uecker, M.; Hosseini, A. S. A.; Schmidt, J. Real‐Time MRI With Deep Learning for Efficient Evaluation of Neuromuscular Breathing Impairment.
MedComm 2026, 7 (3), e70579. https://onlinelibrary.wiley.com/doi/10.1002/mco2.70579 - Brinke, K. A. D.; Borsch, L.-M.; Klose, C.; Zschüntzsch, J.; Vinhoven, L.; Nietert, M.; Justus Lattau, S. S.; Fitzner, D. Plasma Lipidomic Patterns Associated with Disease Activity in Chronic Inflammatory Demyelinating Polyradiculoneuropathy (LIPID-CIDP). Journal of Lipid Research 2025, 100903. https://doi.org/10.1016/j.jlr.2025.100903.
- Rodriguez Gonzalez, C.; Basílio-Queirós, D.; Neehus, A.-L.; Merkert, S.; Tschritter, D.; Ünal, S.; Hegermann, J.; Mörgelin, M.; Bustamante, J.; Nietert, M. M.; Martin, U.; Tümmler, B.; Munder, A.; Lachmann, N. Human CFTR Deficient iPSC-Macrophages Reveal Impaired Functional and Transcriptomic Response upon Pseudomonas Aeruginosa Infection. Front. Immunol. 2024, 15. https://doi.org/10.3389/fimmu.2024.1397886.
- Wei, W.; Lattau, S. S. J.; Xin, W.; Pan, Y.; Tatenhorst, L.; Zhang, L.; Graf, I.; Kuang, Y.; Zheng, X.; Hao, Z.; Popa-Wagner, A.; Gerner, S. T.; Huber, S.; Nietert, M.; Klose, C.; Kilic, E.; Hermann, D. M.; Bähr, M.; Huttner, H. B.; Liu, H.; Fitzner, D.; Doeppner, T. R. Dynamic Brain Lipid Profiles Modulate Microglial Lipid Droplet Accumulation and Inflammation Under Ischemic Conditions in Mice. Advanced Science 2024, n/a (n/a), 2306863. https://doi.org/10.1002/advs.202306863.
- Bachanek, S.; Wuerzberg, P.; Biggemann, L.; Janssen, T. Y.; Nietert, M.; Lotz, J.; Zeuschner, P.; Maßmann, A.; Uhlig, A.; Uhlig, J. Renal Tumor Segmentation, Visualization, and Segmentation Confidence Using Ensembles of Neural Networks in Patients Undergoing Surgical Resection. Eur Radiol 2024. https://doi.org/10.1007/s00330-024-11026-6.
- Lattau, S. S. J.; Borsch, L.-M.; Auf Dem Brinke, K.; Klose, C.; Vinhoven, L.; Nietert, M.; Fitzner, D. Plasma Lipidomic Profiling Using Mass Spectrometry for Multiple Sclerosis Diagnosis and Disease Activity Stratification (LipidMS). IJMS 2024, 25 (5), 2483. https://doi.org/10.3390/ijms25052483.
- Pallenberg S. T., Held I., Dopfer C., Minso R., Nietert M.M., Hansen G., Tümmler B., and Dittrich A.M..
Differential Effects of ELX/TEZ/IVA on Organ-Specific CFTR Function in Two Patients with the Rare CFTR Splice Mutations c.273+1G>A and c.165-2A>G.
Frontiers in Pharmacology, Sec. Pharmacology of Ion Channels and Channelopathies, 14 (March 15, 2023). https://doi.org/10.3389/fphar.2023.1153656. - Vinhoven L.; Stanke F.; Hafkemeyer S.; Nietert M.:
Complementary Dual Approach for In Silico Target Identification of Potential Pharmaceutical Compounds in Cystic Fibrosis.
International Journal of Molecular Sciences, Section Biochemistry, Special Issue: Small Molecule Drug Design and Research (accepted October 2022). https://www.mdpi.com/1422-0067/23/20/12351 - Voskamp M.; Vinhoven L.; Stanke F.; Hafkemeyer S.; Nietert M.:
Integrating Text Mining into the Curation of Disease Maps.
Biomolecules 12, no. 9 (September 2022): 1278. https://doi.org/10.3390/biom12091278. - Nietert M.; Vinhoven L.; Auer F.; Hafkemeyer S.; Stanke F.:
Comprehensive analysis of chemical structures that have been tested as CFTR activating substances in a publicly available database CandActCFTR
Frontiers in Pharmacology,2021. https://www.frontiersin.org/articles/10.3389/fphar.2021.689205 - Pallenberg S., T.; Junge S.; Ringshausen C.; F., Sauer-Heilborn A.; Hansen G.; Dittrich A., M.; Tümmler B.; Nietert M.:
CFTR modulation with elexacaftor-tezacaftor-ivacaftor in people with cystic fibrosis assessed by the β-adrenergic sweat rate assay
Journal of Cystic Fibrosis,2021. https://doi.org/10.1016/j.jcf.2021.10.005 - Vinhoven L.; Voskamp M.; Nietert M.M.
Mapping Compound Databases to Disease Maps—A MINERVA Plugin for CandActBase.
J. Pers. Med. 2021, 11, 1072. doi.org/10.3390/jpm11111072 - Vinhoven L.; Stanke F.; Hafkemeyer S.; Nietert M.M.
CFTR Lifecycle Map—A Systems Medicine Model of CFTR Maturation to Predict Possible Active Compound Combinations.
Int. J. Mol. Sci. 2021, 22(14), 7590; https://doi.org/10.3390/ijms22147590; accepted Jul 2021. - Bleckmann A, Kirchner B, Nietert M, Peeck M, Balkenhol M, Egert D, Rohde TV, Beißbarth T, Pukrop T.
Impact of pre-OP independence in patients with limited brain metastases on long-term survival.
BMC Cancer. 2020 Oct 8;20(1):973. PMID: 33032552 doi.org/10.1186/s12885-020-07459-z. - Jo P, Bernhardt M, Nietert M, König A, Azizian A, Schirmer MA, Grade M, Kitz J, Reuter-Jessen K, Ghadimi M, Ströbel P, Schildhaus HU, Gaedcke J.
KRAS mutation status concordance between the primary tumor and the corresponding metastasis in patients with rectal cancer.
PLoS One. 2020 Oct 1;15(10):e0239806. eCollection 2020. PMID: 33002027 doi.org/10.1371/journal.pone.0239806. - Uhlig J, Biggemann L, Nietert MM, Beißbarth T, Lotz J, Kim HS, Trojan L, Uhlig A.
Discriminating malignant and benign clinical T1 renal masses on computed tomography: A pragmatic radiomics and machine learning approach.
Medicine (Baltimore). 2020 Apr;99(16):e19725. PMID: 32311963 doi.org/10.1097/MD.0000000000019725. - Jo P, Kesruek H, Nietert M, Sahlmann C, Gaedcke J, Ghadimi M, Sperling J.
Inzidenz und Prädiktive Faktoren des Bilateralen Papillären Schilddrüsenkarzinoms.
Zentralblatt für Chirurgie. 2018 Aug;143(4):361-366. Epub 2018 Aug 22, PMID: 30134494 doi.org/10.1055/a-0651-0878. - Lowes M, Kleiss M, Lueck R, Detken S, Koenig A, Nietert M, Beissbarth T, Stanek K, Langer C, Ghadimi M, Conradi LC, Homayounfar K.
The utilization of multidisciplinary tumor boards (MDT) in clinical routine: results of a health care research study focusing on patients with metastasized colorectal cancer.
International journal of colorectal disease. 2017; PMID: 28779354, PMCID: PMC5596058, dx.doi.org/10.1007%2Fs00384-017-2871-z - Linke F, Harenberg M, Nietert M, Zaunig S, von Bonin F, Arlt A, Szczepanowski M, Weich HA, Lutz S, Dullin C, Janovská P, Krafčíková M, Trantírek L, Ovesná P, Klapper W, Beissbarth T, Alves F, Bryja V, Trümper L, Wilting J, Kube D.
Microenvironmental interactions between endothelial and lymphoma cells: a role for the canonical WNT pathway in Hodgkin lymphoma.
Leukemia. 2017; 31(2):361-372. PMID: 27535218 doi.org/10.1038/leu.2016.232 - Linke F, Zaunig S, Nietert M, von Bonin F, Lutz S, Dullin C, Janovská P, Beissbarth T, Alves F, Klapper W, Bryja V, Pukrop T, Trümper L, Wilting J, Kube D.
WNT5A: a motility-promoting factor in Hodgkin lymphoma.
Oncogene. 2017; 36(1):13-23. PMID: 27270428 doi.org/10.1038/onc.2016.183 - Rühlmann F, Nietert M, Sprenger T, Wolff HA, Homayounfar K, Middel P, Bohnenberger H, Beissbarth T, Ghadimi BM, Liersch T, Conradi LC.
The Prognostic Value of Tyrosine Kinase SRC Expression in Locally Advanced Rectal Cancer.
Journal of Cancer. 2017; 8(7):1229-1237. PMID: 28607598, PMCID: PMC5463438 dx.doi.org/10.7150/jca.16980 - Jo P, Nietert M, Gusky L, Kitz J, Conradi LC, Müller-Dornieden A, Schüler P, Wolff HA, Rüschoff J, Ströbel P, Grade M, Liersch T, Beißbarth T, Ghadimi MB, Sax U, Gaedcke J.
Neoadjuvant Therapy in Rectal Cancer - Biobanking of Preoperative Tumor Biopsies.
Scientifc reports. 2016; 6:35589. PMID: 27752113, PMCID: PMC5067705 doi.org/10.1038/srep35589 - Styczen H, Nagelmeier I, Beissbarth T, Nietert M, Homayounfar K, Sprenger T, Boczek U, Stanek K, Kitz J, Wolff HA, Ghadimi BM, Middel P, Liersch T, RüschoffJ, Conradi LC.
HER-2 and HER-3 expression in liver metastases of patients with colorectal cancer.
Oncotarget. 2015; 6(17):15065-76. PMID: 25915155, PMCID: PMC4558136 dx.doi.org/10.18632%2Foncotarget.3527 - von der Heyde S, Wagner S, Czerny A, Nietert M, Ludewig F, Salinas-Riester G, Arlt D, Beißbarth T.
mRNA profling reveals determinants of trastuzumab efciency in HER2-positive breast cancer.
Public Library of Science One. 2015; 10(2):e0117818. PMID: 25710561, PMCID: PMC4339844 dx.doi.org/10.1371%2Fjournal.pone.0117818 - Conradi LC, Styczen H, Sprenger T, Wolff HA, Rödel C, Nietert M, Homayounfar K, Gaedcke J, Kitz J, Talaulicar R, Becker H, Ghadimi M, Middel P, Beissbarth T, Rüschoff J, Liersch T.
Frequency of HER-2 positivity in rectal cancer and Prognosis.
The American journal of surgical pathology. 2013; 37(4):522-31. PMID: 23282976 doi.org/10.1097/pas.0b013e318272ff4d - Tanrikulu Y, Nietert M, Scheffer U, Proschak E, Grabowski K, Schneider P, Weidlich M, Karas M, Göbel M, Schneider G.
Scafold hopping by fuzzy pharmacophores and its application to RNA targets.
Chembiochem : a European journal of chemical biology. 2007; 8(16):1932-6. PMID: 17896338 doi.org/10.1002/cbic.200700195 - Böcker A, Sasse BC, Nietert M, Stark H, Schneider G.
GPCR targeted library design: novel dopamine D3 receptor ligands.
ChemMedChem: Chemistry Enabling Drug Discovery. 2007; 2(7):1000-5. PMID: 17477344 doi.org/10.1002/cmdc.200700067
That might also interest you